Introduction
The population of the microbiota of the human gastrointestinal (GI) tract is
widely diverse and complex with a high population density. All major groups
of organisms are represented in relative amounts that change enormously
from mouth
to anus (see Figure 1 below). The populations are disrupted by infections,
antibiotics, inadequate digestion, and immune system stress.1 While
the microbial mass is comprised predominately of bacteria, a variety
of fungi and protozoa
are also
present. In the colon, there are over 1011 bacterial cells per gram and over
400 different species. These bacterial cells outnumber host cells by at least
a factor of ten. This microbial population has important influences on host
physiological, nutritional, and immunological processes. In fact, this
biomass should more rightly
be considered a rapidly adapting, renewable organ with considerable metabolic
activity and significant influence on human health. Consequently, there is
renewed and growing interest in routine clinical identification of the
types and activities
of gut microbes.2
The normal, healthy balance in microbiota provides colonization resistance
to pathogens. Since anaerobes comprise over 95% of the bacterial population,
their
analysis is of prime importance. Gut microbes might also stimulate immune responses
to prevent conditions such as intestinal dysbiosis (a state of disordered microbial
ecology that causes disease). Specifically, the concept of dysbiosis rests
on the assumption that patterns of intestinal flora, specifically overgrowth
of
some microorganisms found commonly in intestinal flora, have an impact on human
health. Symptoms and conditions thought to be caused or complicated by dysbiosis
include inflammatory bowel diseases, inflammatory or autoimmune disorders,
food allergy, atopic eczema, unexplained fatigue, arthritis, mental/emotional
disorders
in both children and adults, malnutrition, and breast and colon cancers.3,4
Difficulties in Accurately Assessing Microbiota Content
Most studies of microbiota in the human GI tract have used fecal samples. These
do not necessarily represent the populations along the entire GI tract from
stomach to rectum. Conditions and species can alter greatly along this tract
and generally increase from lower to higher population densities as shown
in Figure 1. The stomach and proximal small intestine with highly acid conditions
and rapid flow contain 103 to 105 bacteria per gram of content. These are
predominated by acid tolerant lactobacilli and streptococci bacteria. The
distal small intestine to the ileocecal valve usually ranges to 108 bacteria
per gram. The large intestine generates the highest growth due to longer
residence time, and population densities range from 1010 to 1011 bacteria
per gram. The bacterial metabolism of this region generates a low redox potential
and high amount of short-chain fatty acids.
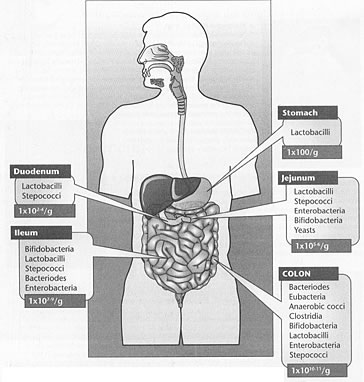
Figure 1: Type and Amount of Bacteria in
Regions of the Gut
For a typical healthy individual, bacterial populations change greating
moving from stomach to stool. The genus or class of predominant organism
is shown inside each box, and the total number of microbes per gram
of intestinal content is shown at the bottom of the box.
Not
only does the total microbiota change throughout the length of the
GI tract, but there are different microenvironments where
these
organisms grow. At least
four microhabitats exist: the intestinal lumen, the unstirred mucus layer
that covers the epithelium, the deeper mucus layer in the crypts
between villi,
and the surface mucosa of the epithelial cells.5,6 Given this
diverse ecological community, the question arises as to how to sample the
various environments
to identify populations of microbes and ultimately understand the host-microbe
interactions. This problem is an extremely difficult one, since any intervention
to obtain a sample potentially disrupts the population. The practice of
fecal sampling should be understood primarily to represent organisms growing
in
the colon. Since more than 98% of fecal bacteria will not grow in oxygen,5 standard
culture techniques miss the majority of organisms present. Conventional Techniques Vs. New Technologies
Conventional bacteriological methods like microscopy, culture, and
identification may be used for the analysis and/or quantification of
the intestinal microbiota.7-9
Limitations of conventional methods include low sensitivities,10 inability
to detect noncultivatable bacteria and unknown species, time-consuming aspects,
and low levels of reproducibility due to the multitude of species to be identified
and quantified. In addition, the large differences in growth rates and growth
requirements of the different species present in the human gut indicate that
quantification by culture is bound to be inaccurate due to restrictions of
growth media choices. To overcome the problems of culture, techniques based
on 16S ribosomal DNA (rDNA) genes were developed.11,12 These include fluorescent
in situ hybridization,13-17 denaturing gradient gel electrophoresis,18,20
and temperature gradient gel electrophoresis. These techniques have high sensitivities,
although they are laborious and technically demanding.
Another problematic issue with present stool analysis procedures is that of
specimen transport. Since growth in culture media requires living organisms,
sample collection must be done using nutrient broth containers to maintain
microbial viability. This allows continued growth of species during transport
and until the sample is actually plated out for culture in the laboratory.
This growth allows for a significant change in the balance of microbes from
that which was present in the patient. Some species will more actively grow
at the expense of others. DNA analysis eliminates this problem by placing the
specimen in formalin vials for transport. This immediately kills all organisms,
freezing the exact balance present at the time of collection. Since PCR identification
is only looking for the genes of the microbiota, living specimens are not necessary.
(The DNA of ingested bacteria is generally degraded, so it is not detected
in the stool specimen.) The more accurate assessment of populations in the
patient's colon allows the clinician to develop the most appropriate
therapy based on the patient's true gut microbiota, resulting in better
clinical results.
Polymerase Chain Reaction (PCR)
One of the most important and profound contributions to molecular biology is
the advent of the polymerase chain reaction (PCR). It is a powerful tool,
enabling us to detect a single genome of an infectious agent in any body
fluid with high accuracy and sensitivity. Many infectious agents that are
missed by routine cultures, serological assays, DNA probes, and Southern
blot hybridizations can be detected by PCR. Therefore, PCR-based tests are
best suited for the clinical and epidemiological investigation of pathogenic
bacteria and viruses. The introduction of PCR in the late 1980s dominated
the clinical market, because it was superior to all previously used culture
techniques and the more recently developed DNA probes and kits. PCR-based
tests are several orders of magnitude more sensitive than those based on
direct hybridization with the DNA probe. PCR does not depend on the ability
of an organism to grow in culture. Furthermore, PCR is fast, sensitive, and
capable of copying a single DNA sequence of a viable or non-viable cell over
a billion times within three to five hours. The sensitivity of the PCR test
is also based on the fact that PCR methodology requires only one to five
cells for detection, whereas a positive culture requires an inoculum equivalent
to about 1000 to 5000 cells. This difference makes PCR the most sensitive
detection method available by several orders of magnitude.21
Advantages of PCR
Amplifications of Target Microbial DNA for Organism Detection:
- Ability
to detect non-viable organisms that are not retrievable by culture
based methods
- Ability to detect and identify organisms that cannot
be cultured or are extremely difficult to grow (e.g., anaerobes)
- More rapid detection and identification of organisms that grow
slowly (e.g., mycobacteria and fungi)
- Ability to detect entire classes
or previously unknown organisms directly in clinical specimens
by using broad range primers
- Ability to quantitate infectious organism
burden for better clinical responsiveness
Laboratories that make the transition to
molecular diagnostics will become an integral part of hospital operations
as they demonstrate
the value of their
improved services. The clinical microbiology laboratory is transitioning
into the molecular age. Through rapid pathogen and antibiotic resistance
identification
and screening tests, rapid molecular diagnostics are playing an increasingly
important role in diagnosing and preventing infections and improving overall
hospital operations. As physicians, pharmacists, and even hospital administrators
demand rapid microbiology results, many laboratories are focusing on being
part of cross-functional implementation teams that not only assure the new
tests are implemented efficiently, but that the results affect real change
for patient management, hospital operations, and laboratory efficacy. Parasitology
Parasitology is yet another field of microbiology to be greatly improved with
molecular technologies. Parasite infections are a major cause of nonviral
diarrhea even in developed countries. Classically, parasites have been identified
by microscopy and enzyme immunoassays.22 In recent studies, molecular techniques
have proven to be more sensitive and specific than classic laboratory methods.22-24
Because Giardia cysts are shed sporadically and the number may vary from
day to day, laboratories have adopted multiple stool collections to help
increase identification rates for all parasite examinations.23 Even with
the advent of antigen detection systems, there has long been uncertainty
in diagnosis when no ova or parasites are found in the stool. Due to the
nearly 100% sensitivity and specificity of DNA analysis, combined with the
need for very low amounts of genomic DNA (as low as 2.5 cells per gram),23
the multiple-day specimen collection, laborious and technically challenging
microscopy, and resulting delays in reporting have been alleviated. With
PCR technology, only one fecal sample is needed for 100% sensitivity and
specificity in parasitology examinations.
Detection of Antibiotic-Resistance Genes
The development of bacterial resistance to antibiotic drugs involves an active
change or mutation in the microbial genome that alters the microbe's
metabolic or structural responsiveness to the mechanism of the drug's
action. This genetic change is passed in the population as cells replicate.
This genetic material can also be passed on to other strains of bacteria
through plasmid sharing. The development of antibiotic resistance is becoming
a serious public health issue, as overuse of antibiotics continually selects
for mutated strains that have developed resistance.
The human intestinal microbiota represent over 400 species. All antibiotic
resistance strategies that bacteria develop are encoded in one or more genes.
These genes are readily shared among and across species and genera and even
among distantly related bacteria. These genes confer resistance to different
classes of drugs, and their sequences are known. Using PCR techniques, an antibiotic
resistance gene in a single organism can be readily detected in large microbial
populations like those found in fecal material.
The knowledge of the presence of a drug-resistant gene may be quite significant
for the clinician when considering treatment of a patient for a pathogen infection.
For example, suppose a pathogen is detected in a stool analysis. An analysis
of the presence of antibiotic resistance genes is also performed on the sample,
and drug sensitivities are then run on the pathogen. It is found to be sensitive
to two antibiotics. But suppose there is also a drug-resistant gene present
in the sample to one of the drugs (a very possible scenario). It would be imperative,
then, that this drug not be used in treating the patient. Otherwise, even though
the pathogen is killed, the other organisms that have the gene conferring resistance
to the drug would thrive relative to other microbes present. This would set
up a potentially dangerous situation where antibiotic resistance is propagated
in the population, because that gene can be readily spread to other organisms
present in the individual.25-27 Knowledge of the presence of antibiotic resistance
genes in fecal specimens, therefore, represents a significant advance in the
treatment of patients and maintenance of health.
Conclusion
DNA analysis technology allows for a significant advancement in understanding
of how GI tract microbiota affect human health. It improves patient care
by giving clinicians greater options and more tools in treating patients.
The increased speed of analysis and improved accuracy offers the potential
for making this the standard method of stool analysis.

Notes
1. Fuller R, Perdigón G. Gut Flora, Nutrition,
Immunity and Health. Oxford; Malden, MA: Blackwell Pub.; 2003.
2. Mackie RI, Sghir A, Gaskins HR. Developmental microbial ecology
of the neonatal gastrointestinal tract. Am
J Clin Nutr. May 1999;69(5):1035S-1045S.
3. Hawrelak JA, Myers SP. The causes of intestinal dysbiosis: a review.
Altern Med Rev. Jun 2004;9(2):180-197.
4. Galland L, Barrie S. Intestinal dysbiosis and the causes of diseases.
J. Advancement Med. 1993;6:67-82.
5. Savage DC. Microbial ecology of the gastrointestinal tract. Annu
Rev Microbiol. 1977;31:107-133.
6. Berg RD. The indigenous gastrointestinal microflora. Trends
Microbiol.
Nov 1996;4(11):430-435.
7. O'Sullivan DJ. Methods of analysis of the intestinal microflora.
In: Tannock GW, ed. Probiotics: A Critical
Review. Wymondham: Horizon
Scientific Press; 1999:23-44.
8. Tannock GW. Analysis of the intestinal microflora: a renaissance.
Antonie Van Leeuwenhoek. Jul-Nov 1999;76(1-4):265-278.
9. Finegold S, Sutter V, Mathisen G. Normal
indigenous intestinal flora.
New York: Academic Press; 1983.
10. Dutta S, Chatterjee A, Dutta P, et al. Sensitivity and performance
characteristics of a direct PCR with stool samples in comparison to
conventional techniques for diagnosis of Shigella and enteroinvasive
Escherichia coli infection in children with acute diarrhoea in Calcutta,
India. J Med Microbiol. Aug 2001;50(8):667-674.
11. Amann RI, Ludwig W, Schleifer KH. Phylogenetic identification and
in situ detection of individual microbial cells without cultivation.
Microbiol Rev. Mar 1995;59(1):143-169.
12. Wilson KH, Blitchington RB. Human colonic biota studied by ribosomal
DNA sequence analysis. Appl Environ Microbiol. Jul 1996;62(7):2273-2278.
13. Franks AH, Harmsen HJ, Raangs GC, Jansen GJ, Schut F, Welling GW.
Variations of bacterial populations in human feces measured by fluorescent
in situ hybridization with group-specific 16S rRNA-targeted oligonucleotide
probes. Appl Environ Microbiol. Sep 1998;64(9):3336-3345.
14. Jansen GJ, Mooibroek M, Idema J, Harmsen HJ, Welling GW, Degener
JE. Rapid identification of bacteria in blood cultures by using fluorescently
labeled oligonucleotide probes. J Clin Microbiol. Feb 2000;38(2):814-817.
15. Langendijk PS, Schut F, Jansen GJ, et al. Quantitative fluorescence
in situ hybridization of Bifidobacterium spp. with genus-specific 16S
rRNA-targeted probes and its application in fecal samples. Appl
Environ Microbiol. Aug 1995;61(8):3069-3075.
16. Muyzer G, Smalla K. Application of denaturing gradient gel electrophoresis
(DGGE) and temperature gradient gel electrophoresis (TGGE) in microbial
ecology. Antonie Van Leeuwenhoek. Jan 1998;73(1):127-141.
17. Welling GW, Elfferich P, Raangs GC, Wildeboer-Veloo AC, Jansen
GJ, Degener JE. 16S ribosomal RNA-targeted oligonucleotide probes for
monitoring of intestinal tract bacteria. Scand
J Gastroenterol Suppl.
1997;222:17-19.
18. Suau A, Bonnet R, Sutren M, et al. Direct analysis of genes encoding
16S rRNA from complex communities reveals many novel molecular species
within the human gut. Appl Environ Microbiol. Nov 1999;65(11):4799-4807.
19. Simpson JM, McCracken VJ, White BA, Gaskins HR, Mackie RI. Application
of denaturant gradient gel electrophoresis for the analysis of the
porcine gastrointestinal microbiota. J Microbiol
Methods. Jun 1999;36(3):167-179.
20. Zoetendal EG, Akkermans AD, De Vos WM. Temperature gradient gel
electrophoresis analysis of 16S rRNA from human fecal samples reveals
stable and host-specific communities of active bacteria. Appl
Environ Microbiol. Oct 1998;64(10):3854-3859.
21. Forbes BA, Sahm DF, Weissfeld AS, Baron EJ. Bailey & Scott's
Diagnostic Microbiology. 10th ed. St. Louis: Mosby; 1998.
22. Verweij JJ, Blange RA, Templeton K, et al. Simultaneous detection
of Entamoeba histolytica, Giardia lamblia, and Cryptosporidium parvum
in fecal samples by using multiplex real-time PCR. J
Clin Microbiol. Mar 2004;42(3):1220-1223.
23. Ghosh S, Debnath A, Sil A, De S, Chattopadhyay DJ, Das P. PCR detection
of Giardia lamblia in stool: targeting intergenic spacer region of
multicopy rRNA gene. Mol Cell Probes. Jun 2000;14(3):181-189.
24. Morgan UM, Pallant L, Dwyer BW, Forbes DA, Rich G, Thompson RC.
Comparison of PCR and microscopy for detection of Cryptosporidium parvum
in human fecal specimens: clinical trial. J
Clin Microbiol. Apr 1998;36(4):995-998.
25. Bergeron MG, Ouellette M. Preventing antibiotic resistance through
rapid genotypic identification of bacteria and of their antibiotic
resistance genes in the clinical microbiology laboratory. J
Clin Microbiol. Aug 1998;36(8):2169-2172.
26. Martineau F, Picard FJ, Grenier L, Roy PH, Ouellette M, Bergeron
MG. Multiplex PCR assays for the detection of clinically relevant antibiotic
resistance genes in staphylococci isolated from patients infected after
cardiac surgery. The ESPRIT Trial. J Antimicrob
Chemother. Oct 2000;46(4):527-534.
27. Martineau F, Picard FJ, Lansac N, et al. Correlation between the
resistance genotype determined by multiplex PCR assays and the antibiotic
susceptibility patterns of Staphylococcus aureus and Staphylococcus
epidermidis. Antimicrob Agents Chemother. Feb 2000;44(2):231-238.
|



![]()
![]()
![]()


